Visualizing the Immune Microenvironment in Human Cancers
Single cell technologies provide rich information about the gene expression programs of immune cells in human cancers, but critical spatial information on cell – cell interactions is lost when tumors are dissociated into single cell suspensions for scRNA-seq or high-dimensional flow cytometry. We have therefore implemented the CODEX platform which enables imaging of human tumors with very large panels of DNA-barcoded antibodies. The major strength of this technology platform is that it enables imaging of large tumor areas and also provides significant flexibility in experimental design. We are using this technology to investigate major immune resistance pathways we have identified in mechanistic studies, such as the integrin avb6 – TGFb – SOX4 pathway.
Single cell technologies provide rich information about the gene expression programs of immune cells in human cancers, but critical spatial information on cell – cell interactions is lost when tumors are dissociated into single cell suspensions for scRNA-seq or high-dimensional flow cytometry. We have therefore implemented the CODEX platform which enables imaging of human tumors with very large panels of DNA-barcoded antibodies. The major strength of this technology platform is that it enables imaging of large tumor areas and also provides significant flexibility in experimental design. We are using this technology to investigate major immune resistance pathways we have identified in mechanistic studies, such as the integrin avb6 – TGFb – SOX4 pathway.
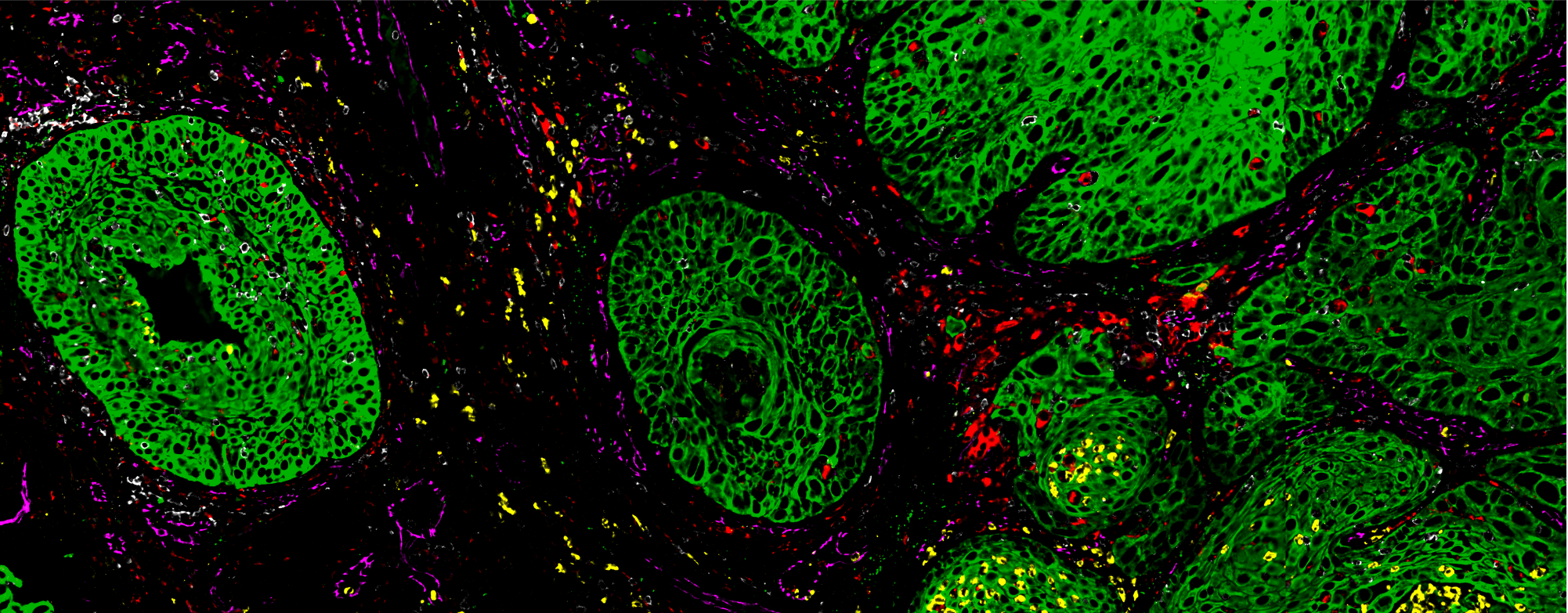
CODEX analysis of human squamous cell carcinoma of the oral cavity: cancer cells (green, pancytokeratin), macrophages (red, CD68), granulocytes (yellow, CD66b), blood vessels (magenta, CD31).
The cancer immunology literature frequently describes tumors as ‘hot’ or ‘cold’. However, preliminary data demonstrate striking regional differences within the same tumor in the degree of immune cell infiltration. We are investigating the molecular mechanisms that determine such striking regional differences because these may be important for resistance to immunotherapy.
We will use this technology in combination with single cell data to study cell – cell communication in the tumor microenvironment and investigate resistance pathways to cancer immunotherapy.
We will use this technology in combination with single cell data to study cell – cell communication in the tumor microenvironment and investigate resistance pathways to cancer immunotherapy.
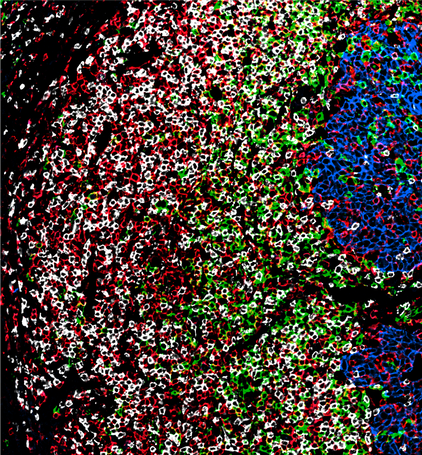
CODEX analysis of highly immune-infiltrated region in a human squamous cell carcinoma of the oral cavity: cancer cells (blue, E-cadherin), CD8 T cells (white, CD8), CD4 T cells (red, CD4), B cells (green, CD20).